Subarea 3: Genetics and Epigenetics of Aging
The focus of Subarea 3 is on genetic and epigenetic determinants of life- and health span as well as aging in fish, rodents and humans. This line of research builds on the expertise of the institute in comparative and functional genomics.
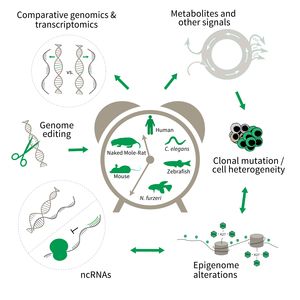
The research is defined by five focus areas:
- Comparative genomics in short- and long-lived models of aging,
- Genomic engineering in N. furzeri,
- Epigenetics of aging,
- Non-coding RNAs in aging, and
- Comparative transcriptomics of aging.
Research focus of Subarea 3.
To uncover causative factors for aging, comparative genomics in short- and long-lived model systems are applied. Functional genomics is used to identify novel pathways contribute to aging of an organism and to validate the functional relevance of genetic and epigenetic changes that occur during aging. Furthermore, genetic risk factors for aging-related diseases are identified and functionally tested. The future development of the Subarea aims to integrate changes in host-microbiota interactions during aging, and how these influence clonal mutation and epigenetic alterations through metabolites and other signals.
Publications
(since 2016)
2017
- Evolution and antiviral specificity of interferon-induced Mx proteins of bats against Ebola-, Influenza-, and other RNA viruses.
Fuchs J, Hölzer M, Schilling M, Patzina C, Schoen A, Hoenen T, Zimmer G, Marz M, Weber F, Müller MA, Kochs G
J Virol 2017, 91(15), e00361-17 - microRNA-122 target sites in the hepatitis C virus RNA NS5B coding region and 3' untranslated region: function in replication and influence of RNA secondary structure.
Gerresheim GK, Dünnes N, Nieder-Röhrmann A, Shalamova LA, Fricke M, Hofacker I, Höner Zu Siederdissen C, Marz M, Niepmann M
Cell Mol Life Sci 2017, 74(4), 747-60 - Age-dependent increase of oxidative stress regulates microRNA-29 family preserving cardiac health.
Heid* J, Cencioni* C, Ripa* R, Baumgart* M, Atlante S, Milano G, Scopece A, Kuenne C, Guenther S, Azzimato V, Farsetti A, Rossi G, Braun T, Pompilio G, Martelli F, Zeiher AM, Cellerino** A, Gaetano** C, Spallotta** F
Sci Rep 2017, 7(1), 16839 * equal contribution, ** co-senior authors - Software Dedicated to Virus Sequence Analysis "Bioinformatics Goes Viral".
Hölzer M, Marz M
In: Advances in Virus Research (edited by Beer M, Höper D) 2017, 99, 233-257, Academic Press, London - High-throughput single-base resolution mapping of RNA 2'-O-methylated residues.
Incarnato D, Anselmi F, Morandi E, Neri F, Maldotti M, Rapelli S, Parlato C, Basile G, Oliviero S
Nucleic Acids Res 2017, 45(3), 1433–1441 published during change of institution - Verification and characterization of an alternative low density lipoprotein receptor-related protein 1 splice variant.
Kolb M, Kurz S, Schäfer A, Huse K, Dietz A, Wichmann G, Birkenmeier G
PLoS One 2017, 12(6), e0180354 - The anti-tumorigenic activity of A2M-A lesson from the naked mole-rat.
Kurz S, Thieme R, Amberg R, Groth M, Jahnke HG, Pieroh P, Horn LC, Kolb M, Huse K, Platzer M, Volke D, Dehghani F, Buzdin A, Engel K, Robitzki A, Hoffmann R, Gockel I, Birkenmeier G
PLoS One 2017, 12(12), e0189514 - A chromosome conformation capture ordered sequence of the barley genome.
Mascher M, Gundlach H, Himmelbach A, Beier S, Twardziok SO, Wicker T, Radchuk V, Dockter C, Hedley PE, Russell J, Bayer M, Ramsay L, Liu H, Haberer G, Zhang XQ, Zhang Q, Barrero RA, Li L, Taudien S, Groth M, Felder M, Hastie A, Šimková H, Staňková H, Vrána J, Chan S, Muñoz-Amatriaín M, Ounit R, Wanamaker S, Bolser D, Colmsee C, Schmutzer T, Aliyeva-Schnorr L, Grasso S, Tanskanen J, Chailyan A, Sampath D, Heavens D, Clissold L, Cao S, Chapman B, Dai F, Han Y, Li H, Li X, Lin C, McCooke JK, Tan C, Wang P, Wang S, Yin S, Zhou G, Poland JA, Bellgard MI, Borisjuk L, Houben A, Doležel J, Ayling S, Lonardi S, Kersey P, Langridge P, Muehlbauer GJ, Clark MD, Caccamo M, Schulman AH, Mayer KFX, Platzer M, Close TJ, Scholz U, Hansson M, Zhang G, Braumann I, Spannagl M, Li C, Waugh R, Stein N
Nature 2017, 544(7651), 427-33 - The Wilms tumor protein Wt1 contributes to female fertility by regulating oviductal proteostasis.
Nathan A, Reinhardt P, Kruspe D, Jörß T, Groth M, Nolte H, Habenicht A, Herrmann J, Holschbach V, Toth B, Krüger M, Wang ZQ, Platzer M, Englert C
Hum Mol Genet 2017, 26(9), 1694-705 - Die Bibliothek im Körper
Neri F
GIT Labor-Fachzeitschrift 2017, 8, 14-7