Subarea 1: Stem Cell Aging
The individual research groups within Subarea 1 investigate the causes and consequences of stem cell aging. The research work spans from basic model organisms over genetic mouse models up to humanized mouse models engrafted with human stem cells.
According to the FLI, with the closure of two groups since 2016 the representation of invertebrate models of stem cell research was reduced in Subarea 1. The institute presumes that the recruitment of new groups should fill this gap.
The research is defined by four focus areas:
- Cell-intrinsic mechanisms limiting the function of aging stem and progenitor cells,
- Aging-associated alterations of stem cell niches and the systemic environment,
- Mechanisms of clonal selection and epigenetic drifts in stem cell aging, and
- Microbiota- and metabolism-induced impairments in stem cell function during aging (in context of the new focus area Microbiota and Aging currently being built up within Subarea 2).
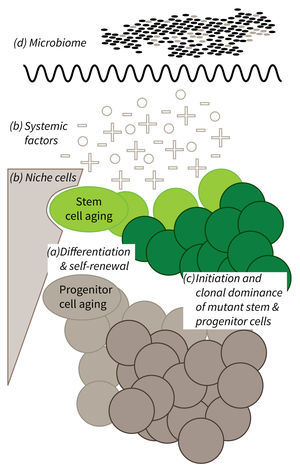
Research focus of Subarea 1.
a) It is currently not well understood what mechanisms impair cellular functions in aging. b) The relative contribution of niche cells and systemic acting factors on stem cell aging have yet to be determined in different tissues. c) Clonal expansion of mutant cells associates with disease development in aging humans. Mechanistically, the process remains poorly understood. Changes in color intensity depict clonal dominance originating from stem (green) or progenitor cells (gray). d) Emerging evidences indicate that aging associated alter ations in microbiota influence stem cell function and vice versa.
Publications
(since 2016)
2025
- How I diagnose and treat patients in the pre-fibrotic phase of primary myelofibrosis (pre-PMF) - practical approaches of a German expert panel discussion in 2024.
Griesshammer M, Al-Ali HK, Eckardt JN, Fiegl M, Göthert J, Jentsch-Ullrich K, Koschmieder S, Kvasnicka HM, Reiter A, Schmidt B, Heidel FH
Ann Hematol 2025, 104(1), 295-306 - Prediction of resistance to hydroxyurea therapy in patients with polycythemia vera: a machine learning study (PV-AIM) validated in a prospective interventional phase IV trial (HU-F-AIM).
Heidel* FH, De Stefano V, Zaiss M, Kisro J, Gückel E, Großer S, Zuurman MW, Manz K, Bryan K, Afsharinejad A, Griesshammer M, Kiladjian JJ
Leukemia 2025, 39(7), 1692-701 * corresponding author - Novel translational mouse models of metabolic dysfunction-associated steatotic liver disease comparable to human MASLD with severe obesity.
Hupa-Breier KL, Schenk H, Campos-Murguia A, Wellhöner F, Heidrich B, Dywicki J, Hartleben B, Böker C, Mall J, Terkamp C, Wilkens L, Becker F, Rudolph KL, Manns MP, Mederacke YS, Marhenke S, Redeker H, Lieber M, Iordanidis K, Taubert R, Wedemeyer H, Noyan F, Hardtke-Wolenski M, Jaeckel E
Mol Metab 2025, 93, 102104 - Cell-free DNA for detection and monitoring of extramedullary AML relapse.
Hupe HC, Wienecke CP, Bartels S, Schipper E, Leßmann J, Lasch A, Bader M, Gabdoulline R, Neugebohren M, Dammann E, Kreipe HH, Lehmann U, Bergmann AK, Di Donato N, Stadler M, Eder M, Ganser A, Heidel FH, Thol F, Heuser M
Hemasphere 2025, 9(3), e70097 - RelB and C/EBPα critically regulate the development of Peyer's patch mononuclear phagocytes.
Kanaya T, Jinnohara T, Sakakibara S, Tachibana N, Sasaki T, Kato T, Riemann M, Jin J, Shiroguchi K, Kawakami E, Ohno H
Mucosal Immunol 2025, 18(1), 151-61 - The histone-methyltransferase DOT1L cooperates with LSD1 to control cell division in blast-phase MPN.
Kapahnke K, Plenge T, Klaus T, Gupta MK, Anand D, Önder TT, Perner B, Schnöder TM, Thol FR, Damm F, Heidel FH, Perner F
Leukemia 2025, 39(10), 2406-18 - The evolving landscape of epigenetic target molecules and therapies in myeloid cancers: focus on acute myeloid leukemia and myeloproliferative neoplasms.
Kühn MWM, Pemmaraju N, Heidel FH
Leukemia 2025, 39(8), 1824-37 - Pathogenesis and management of high molecular risk myeloproliferative neoplasms.
Ling VY, Heidel FH, Bywater MJ
Haematologica 2025, 110(4), 863-76 - Infection and herbicide exposure implicate c-Abl kinase in α-Synuclein Ser129 phosphorylation
Marzieh E, Zeyang S, Alvaro QO, Gesa R, Mahdi R, David H, Saskia FE, Thomas FM
bioRxiv 2025, https://doi.org/10.1101/2025.01. - Prognostic impact of clonal representation of myelodysplasia-related gene mutations in acute myeloid leukemia.
Mecklenbrauck R, Borchert N, Gabdoulline R, Poll P, Funke C, Brandes M, Dallmann LK, Fiedler W, Krauter J, Trummer A, Hertenstein B, Müller M, Lübbert M, Schwalenberg M, Voss A, Di Donato N, Bergmann A, Gaidzik V, Döhner K, Döhner H, Ganser A, Heidel FH, Thol FR, Heuser M
Leukemia 2025, 39(7), 1773-7