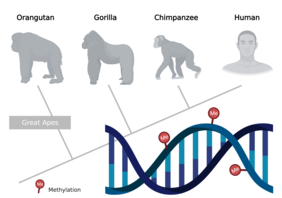
Jena/Los Angeles. Our genetic information is stored in DNA as a sequence of specific base pairings. This is passed on largely unchanged from one generation to the next. But how is it possible that cells differ so massively in form and function, such as in the blood, nerves, skin or teeth, even though they have all the same genetic material (genome)? This is where "epigenetics" comes into play. This primarily refers to chemical changes in our genome that ultimately influence whether and how a cell can access its genetic material. In other words, epigenetic mechanisms determine when which gene is active or inactive. Among the most important epigenetic changes are DNA methylation and the alteration of specific genome-associated proteins (histones). Such changes are also referred to as the epigenome. In contrast to the genome, the epigenome can be rapidly adjusted in response to changes (e.g., environmental factors). Research on the epigenome therefore focuses on the mechanisms that control the activity of genes without altering the base sequence of DNA.
Although the epigenome is attracting increasing attention little is known about how epigenetic changes over time. There is also a lack of systematic investigation strategies. In collaboration with human geneticist Prof. Steve Horvath from the University of California, Los Angeles, USA, researchers from the Jena Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) from the research group “Computational Biology of Aging” and the Core Facility Life Science Computing have shown that classical phylogenetic methods can also be systematically applied to epigenetic data.
Phylogenetic trees based on epigenetic data
In biology, phylogenetic trees have been reconstructed based on sequence-based methods for more than 50 years to comprehensively describe evolutionary events and relationships. "When using epigenetic data, we asked ourselves whether the information they contain enables meaningful evolutionary analysis. Are the signals at all well enough conserved over a longer evolutionary distance to be able to correctly infer phylogenetic relationships from them," says Dr. Arne Sahm, explaining the research approach.
"To gain insights into the function of individual genes and their metabolic pathways, we have used various sequence-based methods in the past, which we have now systematically transferred to the level of DNA methylation," adds Dr. Sahm, first author of the study published in “Molecular Biology and Evolution”. DNA methylation is a direct chemical modification of DNA bases carried out by specialized enzymes in a complicated process. Changes in the start region of genes are particularly consequential: methylation can directly control the activity of a gene.
Epigenetics enables more reliable reconstruction of ancestral relationships
Since our closest relatives - chimpanzees, gorillas and orangutans - differ only slightly from humans in their genomes, the researchers used a dataset of blood samples from great apes for their analyses. "We were able to show that reconstructing phylogenetic trees based on DNA methylation data works very well," reports Prof. Steve Hoffmann, research group leader at FLI and professor of computational biology at Friedrich Schiller University in Jena, Germany. "We were particularly surprised that the ancestral relationships of the great apes can be reconstructed even more reliably with the epigenomic data than with the genomic data. Since methylation is affected by environmental factors or disease, this was not expected."
DNA methylation strongly conserved
The FLI researchers were also able to show that DNA methylation differs only very slightly among the great ape species studied. Thus, genome segments that are highly methylated in humans are generally also altered in the great apes. The same is true for parts of the genome that are poorly methylated or not methylated at all. It is obvious that these epigenomic patterns have remained largely unchanged over millions of years of evolution, i.e., they are strongly conserved. "Here, the resolution of conservation is so high that two equivalent DNA building blocks of different great ape species are on average more similar in their methylation than two directly neighboring DNA building blocks of the same species," Prof. Hoffmann points out.
Insights into tissue-specific phylogenetic information
Unlike the genome, which is nearly identical in all cells of an individual, the epigenome can reflect tissue-specific phylogenetic information. Therefore, comparisons between different species provide the opportunity to gain insight into the evolution of tissues, not just the whole organism. The inclusion of other epigenetic data, such as histone modifications, may be helpful to better understand such processes in the future.
Potential for deeper insights into evolutionary differences and similarities
"The application and further improvement of our developed methods promises new insights into the mechanisms of epigenetic gene regulation and the resulting emergence of new phenotypes," explains Dr. Sahm. The methods could help to identify those parts of the genome that are most likely to play an important role in gene regulation because they have changed little over millions of years. "Conversely, we can also use the method described to identify sections of the epigenome that have changed particularly rapidly in certain species during evolution to study how these epigenetic changes influence the emergence of phenotypes."
Publication
An analysis of methylome evolution in primates. Sahm A, Koch P, Horvath S, Hoffmann S. Mol Biol Evol. 2021, 38(11), 4700-4714. doi: 10.1093/molbev/msab189.
https://academic.oup.com/mbe/article/38/11/4700/6310173
Contact
Dr. Kerstin Wagner
Press and Public Relations
Phone: 03641-656378, email: presse@~@leibniz-fli.de