Englert Research Group
Molecular Genetics:
The Road Map of Aging
Christoph Englert’s Research Group „Molecular Genetics“ lays its research focus on genes regulating organ development and aging.
Molecular Basis of the Urogenital Development
Many "disease" genes in humans play essential roles in the development of specific organs. Examples are the Wilms' tumor suppressor gene Wt1 that, in its mutated form, causes a pediatric kidney cancer and is indispensable for gonad and kidney development in humans and mice. In order to understand how mutations of this gene cause malformations in humans, we are trying to explore the molecular mechanisms by which the respective gene product exerts its function. For this we are employing biochemistry, cell biology as well as animal models.
Signaling Pathways Regulating Aging and Lifespan in Short-Lived Vertebrates
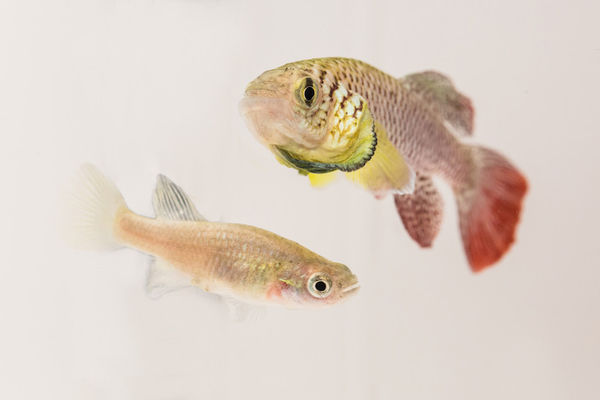
The identification of vertebrate genes, which control aging is hampered by the lifespan of available animal models. Recently, a species of annual fish with an exceptionally short lifespan was described. This species is named Nothobranchius furzeri and has a maximum life expectancy in captivity of just three months. Meanwhile, using CRISPR/Cas9 technology, we are able to switch on and off genes in N. furzeri. We can thereby identify and characterize the genetic programs and biochemical pathways that regulate aging in vertebrates.

Regeneration of Organs
The regenerative capability of human organs differs a lot. While blood cells and skin cells own a high regenerative potential, neurons or kidney cells can regenerate only barely. In contrast, almost all organs of fish and amphibians have a high regenerative potential. We mainly use the zebrafish as animal model to analyze the regeneration processes of caudal fins and kidneys. We are especially interested in understanding how age impacts the regenerative capacity and why the regenerative potential is so different among different species. The ultimate goal is to contribute to increase the regenerative potential e.g. of the human kidney.
Contact

Christoph Englert
Group Leader
+49 3641 65-6042
christoph.englert@~@leibniz-fli.de
Ramona Taubert
Assistance
+49 3641 65-6336
ramona.taubert@~@leibniz-fli.de
Downloads
- CV_CE.pdf98 KB
Selected Publications
- Neuron-specific inactivation of Wt1 alters locomotion in mice and changes interneuron composition in the spinal cord
Schnerwitzki D, Perry S, Ivanova A, Viegas Caixeta F, Cramer P, Guenther S, Weber K, Tafreshiha A, Becker L, Vargas Panesso IL, Klopstock T, Hrabe de Angelis M, Schmidt M, Kullander K, Englert C
Life Science Alliance 2018, 1(4), e201800106 - The Wilms tumor protein Wt1 contributes to female fertility by regulating oviductal proteostasis.
Nathan A, Reinhardt P, Kruspe D, Jörß T, Groth M, Nolte H, Habenicht A, Herrmann J, Holschbach V, Toth B, Krüger M, Wang ZQ, Platzer M, Englert C
Hum Mol Genet 2017, 26(9), 1694-705 - Insights into Sex Chromosome Evolution and Aging from the Genome of a Short-Lived Fish.
Reichwald* K, Petzold* A, Koch* P, Downie* BR, Hartmann* N, Pietsch S, Baumgart M, Chalopin D, Felder M, Bens M, Sahm A, Szafranski K, Taudien S, Groth M, Arisi I, Weise A, Bhatt SS, Sharma V, Kraus JM, Schmid F, Priebe S, Liehr T, Görlach M, Than ME, Hiller M, Kestler HA, Volff JN, Schartl M, Cellerino** A, Englert** C, Platzer** M
Cell 2015, 163(6), 1527-38 * equal contribution, ** co-senior authors, featured in Cell Preview by Vanisha Lakhina and Coleen T. Murphy: Genome Sequencing Fishes out Longevity Genes. Cell. 2015 163(6):1312-3,Nature News by Ewen Callaway: Short-lived fish may hold clues to human age - Integration of Cistromic and Transcriptomic Analyses Identifies Nphs2, Mafb, and Magi2 as Wilms' Tumor 1 Target Genes in Podocyte Differentiation and Maintenance.
Dong L, Pietsch S, Tan Z, Perner B, Sierig R, Kruspe D, Groth M, Witzgall R, Gröne HJ, Platzer M, Englert C
J Am Soc Nephrol 2015, 26(9), 2118-28 - Mitochondrial DNA copy number and function decrease with age in the short-lived fish Nothobranchius furzeri.
Hartmann N, Reichwald K, Wittig I, Dröse S, Schmeisser S, Lück C, Hahn C, Graf M, Gausmann U, Terzibasi E, Cellerino A, Ristow M, Brandt U, Platzer M, Englert C
Aging Cell 2011, 10(5), 824-31