Englert Research Group
Project Biochemistry
Research of the project group Biochemistry focuses on DNA replication and maintenance of genomic integrity in aging and age-related diseases. We study mechanisms of DNA replication, repair and damage response, and the consequences of dysfunction of these processes.
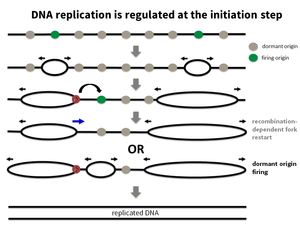
DNA replication, the duplication of the genetic information is a fundamental process in all living cells. Accurate and complete DNA replication is a prerequisite of successful cell division and growth. Therefore, not surprisingly, DNA replication has evolved as a process with amazing plasticity and fidelity. It is further supported by many DNA repair processes and DNA damage response that repair DNA damages in an optimal way. Nevertheless, over time errors during DNA replication slowly accumulate and lead to cell and tissue malfunctions that underlie aging and cancer. This is a particular problem for large and long-living organisms such as humans.
Our research addresses several aspects of DNA replication, repair and damage response:

AgingComb - Investigation of genome aging using high-throughput analysis of DNA replication and recombination
Numerous studies in recent years point to the importance of progressive damage to the genome during ageing. However, the molecular mechanisms that perturb genome integrity in old age are only partially understood. Within the Leibniz ScienceCampus Regenerative Aging, the research groups of Holger Bierhoff (Friedrich Schiller University Jena), Thomas Liehr, Florian Heidel (both University Hospital Jena) and Helmut Pospiech (Leibniz Institute on Aging) cooperate in this field.
The cooperative project AgingComb sheds light on two central processes of DNA metabolism that affect the integrity of the genome: replication and recombination. By using the innovative 'Molecular Combing' technique, changes in the genome structure can be documented in a high-throughput process utilizing the Fiber Combing Platform of Genomic Vision. The data sets generated by this technique allow new insights into the stability of the genome and the molecular mechanisms that affect DNA replication and recombination during aging. By combining our expertise from basic research and clinical practice, we intend to generate benefits for clinical applications.
The AgingComb project is funded by the European Union's European Regional Development Fund (ERDF), which is provided by the Thuringian Ministry of Economics, Science and Digital Society.
Contact

Helmut Pospiech
Project Leader
+49 3641 65-6290
helmut.pospiech@~@leibniz-fli.de
How do hereditary mutations in genes of the BRCA pathway lead to early onset of breast cancer? And how does the imbalance between DNA replication and recombination contribute to genomic instability?
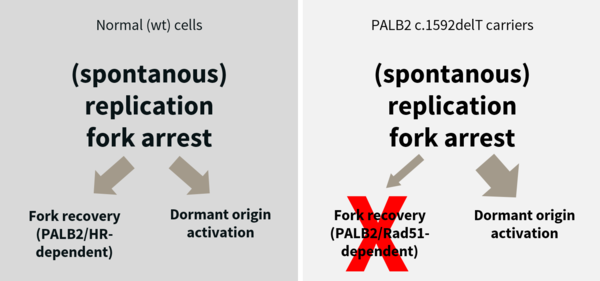
Breast cancer is the most common malignancy in women in industrial countries. Heterozygous mutations in several genes increase the risk and decrease the age of onset to develop breast cancer. Most of the prominent breast cancer genes are involved in the BRCA pathway that regulates homologous recombination after DNA double strand breaks and during DNA replication. We have previously shown in Finnish carriers of a PALB2 mutation that already loss of one of the two copies of this genes leads to a replication stress phenotype as well as defects in the maintenance of cell cycle checkpoints that together lead to genomic instability and is thereby an early driver of cancer (Nikkilä et al., 2013; Obermeier et al., 2016). In collaboration with German and international partners we study the functional consequences of several other hereditary breast cancer mutations. In a DFG-funded project, we study specifically, what are the effects of reduced dose of factors involved in the BRCA pathway on DNA replication, homologous recombination, DNA damage response and genomic stability.
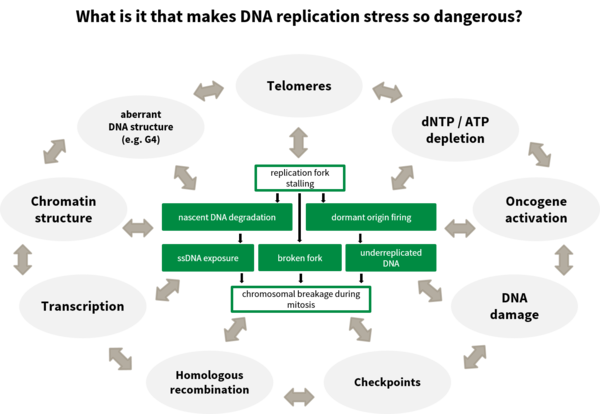
How does DNA replication stress promote genome destabilisation?
DNA replication stress has been identified as a key factor leading to genomic instability and cancer. The number of DNA replication rounds of stem cells in the body and thus the age represent a main determinant of the risk to acquire cancer. The causes of DNA replication stress are manifold as are the phenotypes associated. We have looked at various causes of DNA replication stress such as disturbances of DNA replication (Köhler et al., 2016; Sokka et al., 2018), homologous recombination (Nikkilä et al., 2013; Parplys et al., 2015), DNA damage tolerance (Hampp et al., 2016) as well as metabolic disturbances (Pai et al., 2016), to broaden the understanding of the causes, the phenotypes and the subsequent effects of DNA replication stress.
References:
Hampp, S. et al. (2016). A DNA damage tolerance pathway involving DNA polymerase iota and the tumor suppressor p53 regulates DNA replication fork progression, Proc Natl Acad Sci U S A, 113(30), E4311-9. [PubMed]
Köhler, C. et al. (2016). Cdc45 is limiting for replication initiation in humans, Cell Cycle, 15(7), 974-85. [PubMed]
Nikkilä, J. et al. (2013). Heterozygous mutations in PALB2 cause DNA replication and damage response defects, Nat Commun, 4, 2578. [PubMed]
Pai, G. M. et al. (2016). TSC loss distorts DNA replication programme and sensitises cells to genotoxic stress, Oncotarget, 7(51), 85365-80. [PubMed]
Parplys, A. C. et al. (2015). High levels of RAD51 perturb DNA replication elongation and cause unscheduled origin firing due to impaired CHK1 activation, Cell Cycle, 14(19), 3190-202. [PubMed]
Sokka, M. et al. (2018). The ATR-activation domain of TopBP1 is required for the suppression of origin firing during the S phase, Int J Mol Sci, 19(8), pii: E2376. [PubMed]
What are the limits of mutation rates in human cells and how can mutation rates be control during aging and cancer?
In a more recent line of research we look at the rates of mutations during DNA replication in human cells. We try to define the limits of mutation tolerated by human cells, and the specific characteristics of cells with very high mutation rates. We also look into how mutations accumulate during aging and how organisms with different life expectancy differ in their mutation rates. Finally, we explore new ways how mutation rates can be influenced positively and negatively in the treatment of cancer and the possible alleviation of mutation accumulation during aging.
What are the molecular functions of individual replication proteins?
A longstanding research interest of our project group is the characterization of factors involved in DNA replication and recombination. These include replicative DNA polymerases α, δ and ε (Vaara et al., 2012; Villani et al., 2014; Itkonen et al., 2016), helicase co-factor CDC45 (Szambowska et al., 2014, 2017; Köhler et al., 2016), DNA helicase RecQL4 (Ohlenschläger et al., 2012; Keller et al., 2014) and S100A11 (Foertsch et al., 2016). Recently, we identified a new potential replication factor designated PN70, which is currently functionally characterized in detail.
References:
Foertsch, F. et al. (2016). S100A11 plays a role in homologous recombination and genome maintenance by influencing the persistence of RAD51 in DNA repair foci, Cell Cycle, 15(20), 2766-79. [PubMed]
Itkonen, H. M. et al. (2016). Human DNA polymerase α interacts with mismatch repair proteins MSH2 and MSH6, FEBS Lett, 590(23), 4233-41. [PubMed]
Keller, H. et al. (2014). The intrinsically disordered amino-terminal region of human RecQL4 : multiple DNA-binding domains confer annealing , strand exchange and G4 DNA binding, Nucleic Acids Res, 42(20), 12614-27. [PubMed]
Köhler, C. et al. (2016). Cdc45 is limiting for replication initiation in humans, Cell Cycle, 15(7), 974-85. [PubMed]
Ohlenschläger, O. et al. (2012). The N-terminus of the human RecQL4 helicase is a homeodomain-like DNA interaction motif, Nucleic Acids Res, 40(17), 8309-24. [PubMed]
Szambowska, A. et al. (2014). DNA binding properties of human Cdc45 suggest a function as molecular wedge for DNA unwinding, Nucleic Acids Res, 42(4), 2308-19. [PubMed]
Szambowska, A. et al. (2017). Cdc45-induced loading of human RPA onto single-stranded DNA, Nucleic Acids Res, 45(6), 3217-30. [PubMed]
Vaara, M. et al. (2012). Segregation of replicative DNA polymerases during S phase: DNA polymerase ε, but not DNA polymerases α/δ, are associated with lamins throughout S phase in human cells, J Biol Chem, 287(40), 33327-38. [PubMed]
Villani, G. et al. (2014). Gap-directed translesion DNA synthesis of an abasic site on circular DNA templates by a human replication complex, PLoS One, 9(4), e93908. [PubMed]
Team*
| Name | Phone | Position |
|---|
* incomplete due to Data protection
Find here the Publications of the Project Biochemistry: Publications